Single cell study of intestinal epithelial stem cells with the Nadia
Digest of the publication: Discrete regulation of β-catenin-mediated transcription governs identity of intestinal epithelial stem cells
The gut is an important organ, in charge of absorbing water and nutrients from the food and drinks we ingest. It is also the most highly regenerative organ in the body; regenerating its lining, called epithelium, every five to seven days. This phenomenon relies upon the continuous renewal and proliferation of intestinal epithelial stem cells (IESCs). IESCs reside at the bottom of intestinal crypts, which are invaginations of the epithelium. Crypts neighbour another major type of epithelial structure: villi (singular: villus). Villi are pretty much the opposite of crypts. They are finger-like structures that project into the lumen of the intestine covered with mostly mature absorptive enterocytes and some secretory cells.
Fun fact: villi vastly amplify the surface of the digestive tract. Scientists have calculated that the total surface of the gastro-intestinal tract in healthy adults is between 30 and 40 square meters.
A pre-print recently published by Borrelli et al., looks into the intricate transcriptional mechanisms regulating the maintenance of intestinal homeostasis using a variety of bulk and single cell analysis methods. Specifically, the authors dissected the individual contributions of the C and N-terminal branches of β-catenin. β-catenin is a protein produced in the cytoplasm, which is then translocated into the nucleus, where it acts as a transcriptional activator by forming a transcriptional complex with other factors. Such factors are recruited through interaction with the aforementioned C- and N-terminal branches of the protein. β-catenin exerts its transcriptional control as part of the Wnt pathway, a signal-transducing pathway that regulates gene expression as a response to external stimuli detected through cell surface receptors.
In this study, the authors came to several conclusions including the following:
– The attenuation of the C- and N-terminal branches of β-catenin have contrasting effects on intestinal homeostasis
– The lack of N-terminally-mediated transcriptional outputs leads to transient hyperproliferation of IESCs before complete stem cell loss
– The hyperproliferation of IESCs is driven by cell cycle regulators Myc and E2F
The attenuation of the C- and N-terminal branches of β-catenin have contrasting effects on intestinal homeostasis
To study the effect of attenuating the C- and N-terminal branches of β-catenin, the authors generated transgenic mice expressing such mutated protein in their gut. Importantly, the total lack of functional β-catenin is lethal at the embryonic stage. To overcome this issue, heterozygous mice were created, carrying one mutant and one conditional β-catenin allele. The latter was subjected to deletion following the induction of another transgene coding for the Cre recombinase once mice were 8-12 weeks old. This left the Wnt/β-catenin-mediated transcription under the sole control of the mutated protein and allowed researchers to study the resulting phenotype.
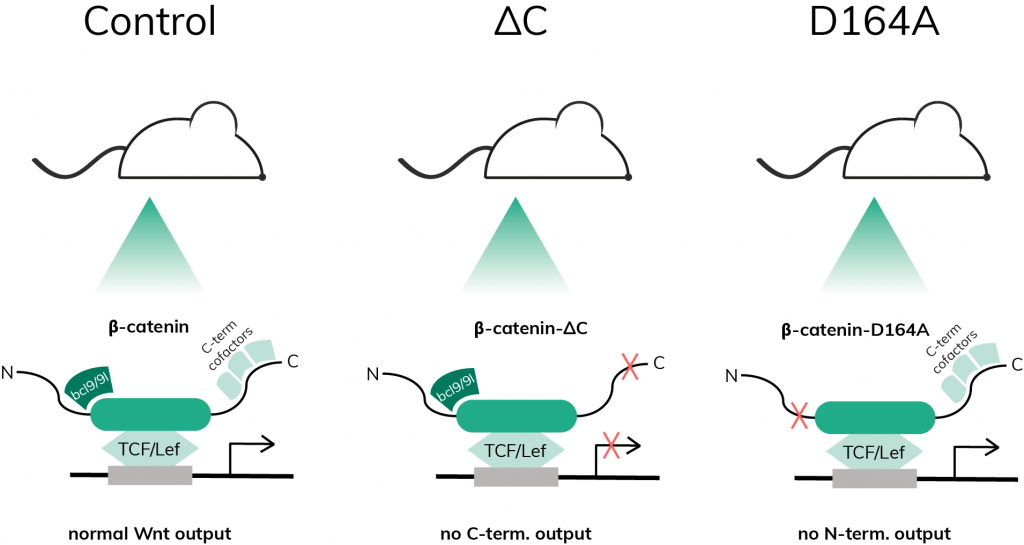
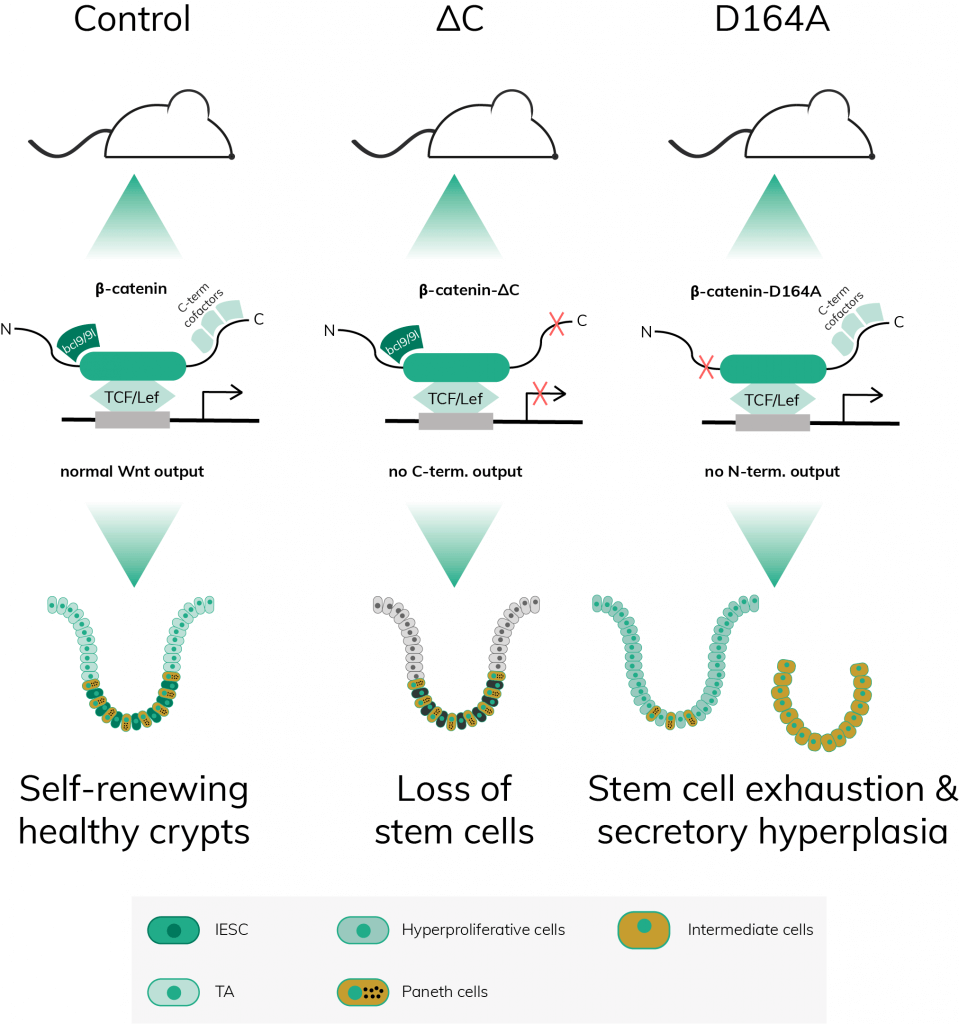
Following this principle, the authors created several lines of transgenic mice, one of them expressing C-terminally mutated β-catenin (referred to as ΔC mice) and another one expressing N-terminally mutated β-catenin (referred to as D164A mice). Four days after injecting the animals with the drug tamoxifen to induce Cre, ΔC mice exhibited atrophic intestinal crypts, a lethal phenotype. On the other hand, D164A mice lived longer and reached humane endpoint at 7d post-induction (pi) when they suffered severe colitis.
The lack of N-terminally-mediated transcriptional outputs leads to transient hyperproliferation of IESCs before complete stem cell loss
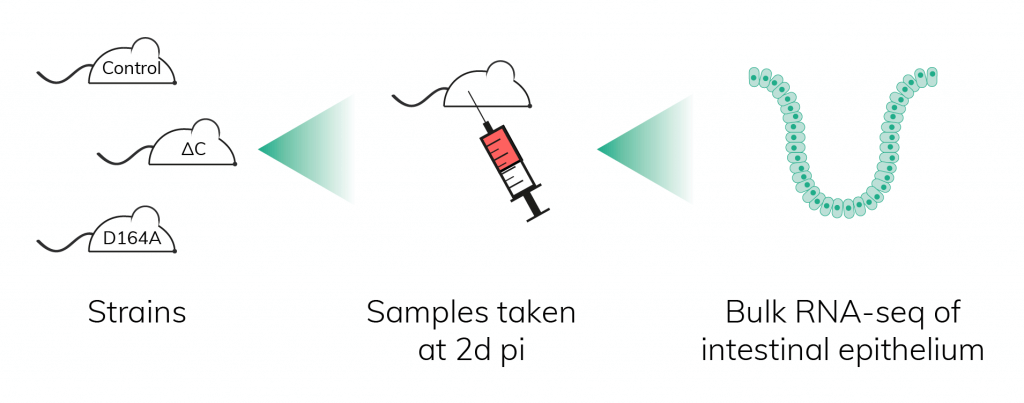
To better understand the cellular and molecular mechanisms responsible for the above dissimilar phenotypes, bulk RNA sequencing was performed with small intestinal epithelium at 2d pi. The differentially expressed genes of ΔC animals were similar to those of full knock-out (KO) mice, i.e. mice in which Cre induction leads to the complete loss of β-catenin. This indicated that C-terminally mutated β-catenin is not able to sustain the level of activity necessary to properly maintain the intestinal epithelium.
In contrast, the differentially expressed genes of D164A mutants only marginally overlapped with those of KO mice. In fact, the results indicated that attenuating the N-terminal branch of β-catenin led to increased proliferation of IESCs. Further investigation showed that this hyperproliferation was only transient. By 4d pi all proliferative markers had disappeared from the transcriptomic profile. Microscopic examination also revealed the presence of unusual, large, secretory cells in the crypts of D164A animals resulting from the mis-differentiation of IESCs.
Taken together these results indicated that neither the C- nor the N-terminally mutated β-catenin are able to substitute the wild-type protein. However, the cellular dynamics leading to lethal tissue damage in both cases are different. While the C-terminal mutation leads to a rapid loss of stem cells, the N-terminal mutation first triggers hyperproliferation of IESCs, followed by aberrant differentiation and eventual stem cell loss.
The hyperproliferation of IESCs is driven by cell cycle regulators Myc and E2F
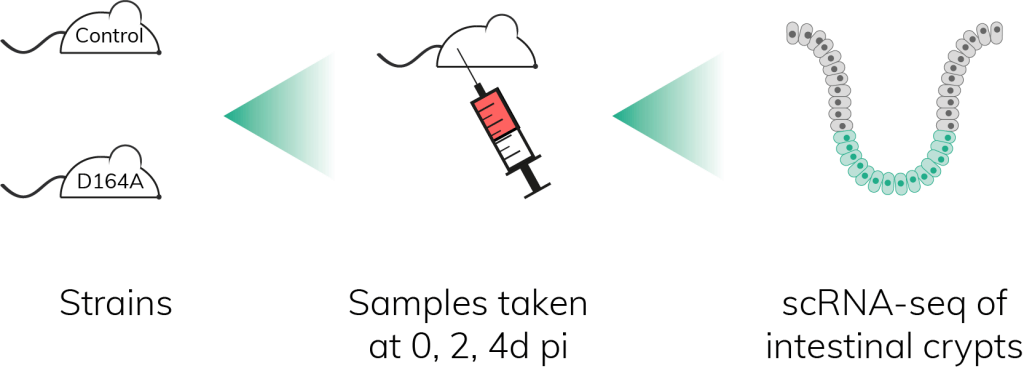
Single-cell RNA sequencing of intestinal crypts was performed on the Nadia Instrument at several time points to study the gene regulatory networks underlying the transient hyperproliferation of IESCs observed in D164A mice. The experiment revealed an enrichment in Myc-dependent Wnt/β-catenin targets at 2d pi in stem cells and early progenitors (SCEPs). Myc proteins are transcription factors involved in the control of the cell cycle – they act on proliferative genes. Other cell cycle genes, including E2F, were upregulated as well. Moreover, in silico comparison of SCEPs’ single-cell datasets against models of IESCs and differentiated progenitors deriving from IESCs (called transit amplifiers, or TAs) showed that hyperproliferative SCEPs exhibited many TA traits. All in all, this showed that impairing β-catenin N-terminal interactions transiently activates an Myc-E2F-driven proliferation of IESCs, which eventually leads to the exhaustion of the self-renewing stem cell pool.
Conclusions
This work demonstrated that while the attenuation of both C- and N-terminal branches of β-catenin leads to lethal intestinal crypt collapse, the cellular mechanisms leading to that point are very different in either case. C-terminally-recruited transcriptional factors appear to act as a straightforward on-off switch. Conversely, N-terminal co-factors were shown to fine-tune Wnt/β-catenin transcriptional outputs, such as those that promote Myc-E2F-driven proliferation, ensuring the maintenance of a self-renewing pool of IESCs. The Wnt/β-catenin pathway and its outputs are therefore tightly regulated within the narrow window of activity required to govern the identity and fate of intestinal epithelial stem cells.
If you are interested in studying human heath using single cell research have a look single cell RNA-Seq with the Nadia platform.
Further reading
Read our previous blogs’ discussing papers involving the Nadia platform
Single cell RNA sequencing of SARS-CoV-2 infected human cell lines
scRNA-Seq in Immunology – A Case Study of the Nadia instrument