NEW Single cell data from the Nadia Instrument
(with real single-cell data examples!)
A lot of people talk about how they want to generate high-quality single cell data that is reproducible and cost-effective. There is a fine art to doing this. It is a smart mix of using the right reagents (that don’t make your research too expensive), following the protocol exactly and using high-quality single cell technology that makes this all possible.
We recently ran a number of single cell runs to prove the Nadia Instrument from Dolomite Bio can do just this. Using the Drop-seq protocol, a 1:1 mixture of HEK293T and 3T3 cells with both 200 cells and 600 cells were used to generate 4 data-sets. The outline of these runs are below:
| Sample | #Cells | NGS reads/cell | Doublet rate | Median Genes | Median UMIs |
| Nadia 200 R1 | 272 | 19757 | 7.3% | 2728 | 6413 |
| Nadia 200 R2 | 185 | 26224 | 6.4% | 3107 | 6299 |
| Nadia 600 R1 | 537 | 121975 | 6.6% | 5525 | 23233 |
| Nadia 600 R2 | 654 | 50024 | 7.8% | 4132 | 11704 |
Full data-sets can be downloaded here.
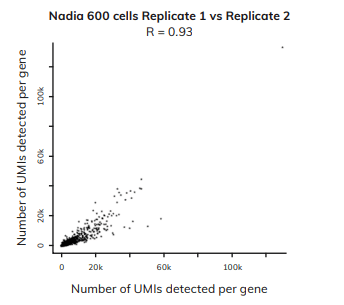
Generating reproducible data
From two of our data-sets, we generated the plot to the left, that outlines the correlation between two datasets from different users generated on different days on the Nadia instrument.
The graph shows that both datasets display high concordance between individual experiments as well as different users. Therefore, demonstrating just how reproducible the data generated using the Nadia Instrument is.
The correlation between all four datasets (showing very high reproducibility!) can be found in more detail in our datasheet.
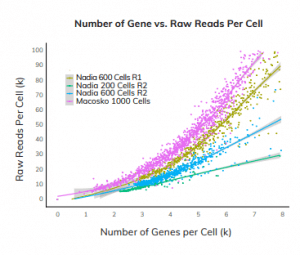
Reducing the cost of single cell profiling
The graph to the left shows the number of detected genes compared to the read depth across all samples, using 3 data-sets and compared to the Macosko data-set.
Efficient gene capture was observed even at low sequencing depth, reducing the cost for NGS analysis. The Nadia Instrument was able to detect gene numbers as high as 2000-3000 at low sequencing depths of 20,000 reads per cell.
The Nadia Instrument is a cost-effective instrument for performing scRNA-Seq.
For a full analysis of our four datasets, please download the datasheet for more details. We are also discussing all our single cell datasets and full analysis on our upcoming webinar – WATCH THE RECORDING HERE.

